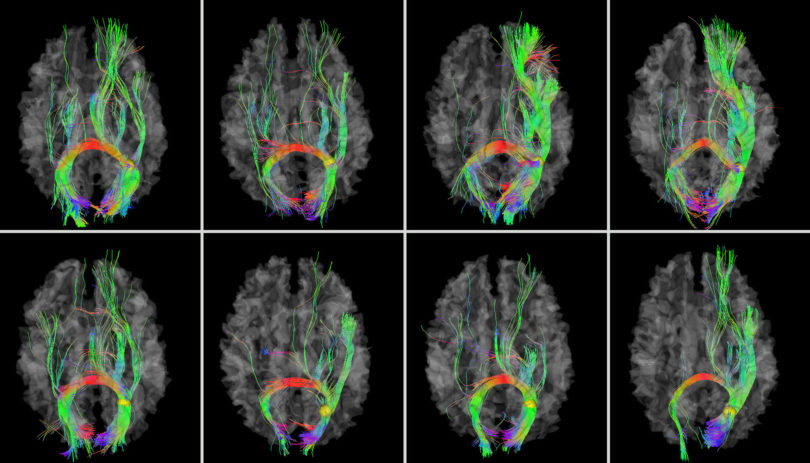
Athens, Ga.- University of Georgia researchers have developed a map of the human brain that shows great promise as a new guide to the inner workings of the body’s most complex and critical organ.
With this map, researchers hope to create a next-generation brain atlas that will be an alternative option to the atlas created by German anatomist Korbinian Brodmann more than 100 years ago, which is still commonly used in clinical and research settings.
Tianming Liu, assistant professor of computer science in the UGA Franklin College of Arts and Sciences, and his students Dajiang Zhu and Kaiming Li identified 358 landmarks throughout the brain related to memory, vision, language, arousal regulation and many other fundamental bodily operations. Their findings were published in the April issue of Cerebral Cortex.
The landmarks were discovered using diffusion tensor imaging, a sophisticated neuroimaging technique that allows scientists to visualize nerve fiber connections throughout the brain. Unlike many other neuroimaging studies, their map does not focus only on one section of the brain but rather the whole cerebral cortex.
“Previously, researchers would examine at most three or four small brain networks,” Liu said. “We want to examine the whole brain connection, and this is the so-called connectome.”
The new map provides a clearer picture of how different areas of the brain are physically connected and how these connections relate to basic brain function. Liu and his team examined hundreds of healthy young adults to establish the landmarks, which they call dense individualized and common connectivity-based cortical landmarks, or DICCCOL.
After extensive testing and comparison, the team determined that these nodes are present in every normal brain, meaning they can be used as a basis of comparison for those with damaged brain tissue or altered brain function.
“DICCCOL is very similar to a GPS system,” Zhu said, “only it’s a GPS map of the human brain.”
Now, thanks in part to a five-year, $1.6 million grant from the National Institutes of Health, Liu and collaborators Xiaoping Hu and Claire Coles at Emory University are preparing to test their brain map by comparing healthy brains with those of children whose brains were damaged by exposure to cocaine while in the womb.
Prenatal cocaine exposure, or PCE, can cause serious damage to brain networks. Because of this, analysis of the damage provides Liu and his team with an excellent opportunity to evaluate the usefulness of their map.
After comparing the PCE brains to those of healthy individuals, they hope to determine the segments of the brain responsible for physical or mental disabilities observed in children exposed to cocaine.
“The PCE brain is disrupted in a systematic way; the whole brain is wrongly wired,” Liu said. “We want to test our map in one of the worst cases, and then we will know if it will work in other cases.”
Once the robustness of their map is established, Liu and his team hope that it may prove useful in the evaluation of many other brain disorders, such as Alzheimer’s disease, Parkinson’s disease or stroke.
“This really is a fundamental technology,” Liu said. “When we establish these DICCCOLS, we can very easily extend this project to other populations, to other brain diseases.”
Liu’s team published their DICCCOL data sets, which includes the source code and diffusion tensor images, at http://dicccol.cs.uga.edu so other researchers may use the findings in their own experiments.
The article, “DICCCOL: Dense Individualized and Common Connectivity-Based Cortical Landmarks,” is available at http://cercor.oxfordjournals.org/content/early/2012/04/05/cercor.bhs072.short?rss=1.